|
|
1.IntroductionIntravascular optical coherence tomography (IVOCT) provides unique high-contrast microscopic resolution imaging of the vasculature.1–12 With regard to atherosclerotic plaques, IVOCT has demonstrated an ability to differentiate lipid, calcium, and fibrous tissue13,14 and to quantify microscopic features such as macrophage content.9 We have previously reported on IVOCT image analysis relating to segmentation, quantification, and visualization of plaques, stents, and other vessel wall components.15–24 We and others are using IVOCT to assess viability of new coronary artery stent designs18–25 and quantitative evaluation of atherosclerotic plaques.15–17 The goal is automated classification and segmentation of plaque types from clinical IVOCT pullbacks. In this report, we focus on computational methods and optimize processing for the reliable determination of tissue optical properties from clinical data. Trained cardiologists and image analysts can identify tissue types (calcified, lipid, and fibrous tissues) with some accuracy and repeatability, as shown in preclinical and clinical studies.4,6,12–14 Many use rules established by Yabushita et al.13 to manually classify tissues. However, modern OCT systems can create over 500 image frames in a single 2.5-s pullback scan, making manual image analysis for research very labor intensive, typically precluding measurements from every image frame. During a demanding clinical procedure, it would be even more difficult to manually analyze hundreds of image frames in clinical decision making. Although IVOCT image quality is outstanding, its limited depth penetration can sometimes confound plaque characterization,6,25–27 posing challenges for manual analysis, especially in the case of lipid or calcified plaques underlying a fibrous cap. Image quality is affected by any residual blood, but this is not a major concern with current IVOCT blood clearing strategies. Additionally, catheter eccentricity during IVOCT image acquisition may alter the appearance of various plaque features and confuse IVOCT image analysts.28 To be successful, an automatic computer classification algorithm will need to use all image information available to the human such as intensity, intensity changes, texture, border sharpness, three-dimensional (3-D) shape characteristics, and, perhaps most importantly, physical optical properties. Several groups have measured tissue optical properties from OCT images in vascular and other tissues using a variety of experimental and theoretical methods. Most use a single-scattering model. We will review reports that measured attenuation and backscattering coefficients in vascular tissues first.29–33 Xu et al.29 used a noncatheter-based approach by imaging transverse, fixed tissue sections of coronary arteries; averaged 400 A-lines over time; applied a single-scattering light model; and performed least squares fitting of log transformed, average A-lines to a linear equation. It was concluded that the calcified and fibrotic plaques had a much lower optical attenuation coefficient, , than lipid plaques, and that lipid and fibrotic plaques had a much higher backscattering coefficient, , than calcified plaques (lipid: , , calcified: , , and fibrotic: , ). Using a catheter-based system and a stationary acquisition without pullback, van Soest et al.30 measured in both in vivo and ex vivo specimens. They corrected for the nonideal imaging system; averaged 15 to 20 A-lines in time; fitted a linear model to log transformed data; and applied an elegant algorithm to determine regions in one dimension corresponding to a single, homogenous tissue type. van Soest et al. argued that cannot be estimated accurately along with using a single magnitude OCT value alone at each point and reported only for calcified (2 to ), fibrotic (2 to ), and lipid (), yielding trends similar to values reported by Xu et al.29 Levitz et al.31 applied a multiple scattering model to OCT images of phantoms and ex vivo aortas and estimated scattering coefficients and anisotropy factors. In other studies, OCT data from both phantoms and ex vivo arterial samples were fit to single and multiple scattering models.32,33 Since temporal averaging of frames from a stationary acquisition is undesirable in the clinical setting, frequency domain multiplexing with improvement as compared with simple linear filtering has been proposed to reduce speckle in single A-lines for optical property measurement.34 As reviewed in Sec. 5, there is variability in reported values between research groups due to methodological differences including parameter estimation approaches. Optical properties have also been estimated in other tissues using OCT. Yang et al.35 determined the optical scattering coefficient, , from unfixed normal and malignant, ex vivo ovarian tissues. They found that the absorption coefficient, , for ovarian tissue is small () and that was a good approximation to . Knu et al.26 determined the optical parameters from OCT images of pig and human skin using a multiple scattering model. Lee et al.36 have studied the performance of single and multiple scattering models using in vitro rat livers and in vivo human skin. The methodologies were similar to those used in blood vessels with 100 to 200 A-lines averaged prior to fitting. In general, it is evident from the literature that a single-scattering model is reasonable and is probably the only statistically appropriate solution for noisy clinical IVOCT images. A goal is to fully automate tissue characterization in 3-D IVOCT clinical pullbacks. Some recent reports use machine learning for plaque classification from both preclinical and clinical IVOCT data.37,38 Ughi et al.38 used estimates from a layer model applied to single A-lines and two-dimensional (2-D) texture and geometric measures as features for classification. Athanasiou et al.37 used 2-D texture and intensity features alone and assumed “islands” of calcified tissue surrounded by other tissue types, which is not necessarily true in many image frames. These reports are encouraging and show that improved 3-D estimation of optical properties could improve the classification. The rationale for our research plan is as follows. Although it is clear that the optical properties of plaques, such as the attenuation coefficients, will be useful discriminatory features for automated tissue classification, extracting optical properties from clinical IVOCT data remains a challenge. Stationary acquisitions29,30 are untenable, as they will require an additional blood clearing operation to sample a single location. Estimation from single, noisy A-lines gives large estimation variance. Hence, our solution is to estimate optical properties from small 3-D volumes of pullback image data and to use advanced parameter estimation approaches which are robust against noise outliers. In addition, we use in vivo clinical data rather than image data from ex vivo samples as is typically used. This avoids typical limitations of ex vivo imaging including potential tissue degradation, no mean and/or pulsating pressure, no cardiac motion, unrealistic temperatures, no blood artifacts, and a straight path pullback rather than actual 3-D coronary geometry. Moreover, in the event of spatially inaccurate, sparse histological sampling, one can make inappropriate “ground truth” determinations using ex vivo data. Instead we use ground truth volumes of interest (VOIs) identified by trained analysts using well-established criteria. Although independent histological verification is desirable, there is no way to obtain it on in vivo clinical data. Moreover, our approach is quite appropriate for optimizing the 3-D parameter estimation methods in this report. Methods should be equally applicable to 3-D cancer imaging. In this report, we correct the nonideal characteristic of OCT imaging systems, use noise reduction filtering, estimate optical parameters from small VOIs rather than single A-lines, reject outliers, identify goodness-of-fit measures suitable for determining the quality of optical property estimates, and determine how well optical properties alone can be used to classify plaque types. Our basic paradigm is to compare estimation results across 311 manually annotated VOIs. 2.Experimental Methods and Catheter CorrectionImages used in this study were selected from the database available at the Cardiovascular Imaging Core Laboratory of the University Hospitals Case Medical Center (Cleveland, Ohio), called the Core Lab. These images were collected on the C7-XR swept-source OCT system (St. Jude Medical Inc., Westford, Massachusetts). It has a swept laser source with a 1310-nm center wavelength, 110-nm wavelength range, 50-kHz sweep rate, 20-mW output power, and coherence length. The pullback speed was and the pullback length was 54 mm. A typical pullback consisted of 271 image frames spaced apart. The test dataset consisted of 35 IVOCT pullbacks of the left anterior descending and the left circumflex coronary arteries and was chosen to be representative of typical OCT scans obtained from patients prior to stent implantation. To more accurately estimate , we need to account for certain characteristics of the IVOCT imaging system. We used methods similar to those of van Soest et al.30 The imaging system response in the presence of tissue attenuation is given below:30 with where is the depth, is the detected intensity, is the initial intensity, is the detected intensity at a depth of , is the attenuation coefficient, is the confocal function with , the position of the beam waist and with , the Rayleigh length, and accounts for the spectral coherence of the source in Fourier-domain OCT with parameters (representing the half-width of the intensity roll off) and (representing the center of the scan).Intact catheters without a guide wire were inserted into a bottle containing water or low concentration intralipid solution. The catheter was firmly held in place using a vice-grip on the catheter outside the bottle. The water image was used to estimate background intensity. A low concentration intralipid (Sigma Aldrich, 20%, emulsion) diluted in water provided an image of scatterers without significant attenuation. Stationary image sequences of 121 image frames and pullback sequences of 271 image frames were obtained. About 100 IVOCT magnitude images were averaged. From the average frame, we averaged 500 A-lines to obtain a very low noise A-line. In intralipid images, we assumed to be low, eliminating the tissue attenuation term []. was estimated from the water images. and were obtained from the interferometer setting and the specifications of the light source, respectively. With these parameters fixed, , , and were estimated by fitting Eq. (1) to average A-line data using nonlinear least squares with the Nelder–Mead simplex optimization. A-line data were truncated to exclude bright reflection from the catheter sheath. Model fits were compared over pullbacks, catheters, and acquisition modes (stationary versus pullback). Similar fits were obtained in each case with less than 3% variation in parameters. Clinical images were corrected using Eq. (1). We subtracted , the baseline intensity, and divided A-line intensities by the term . 3.Image Analysis3.1.Volumes of InterestWe developed specialized software in MATLAB for obtaining user-defined VOIs and for evaluating their optical properties. In the software, pullbacks were loaded and image frames were presented to the expert, an IVOCT image analyst, in both and views. Image data came from a manually analyzed Core Lab study aimed at classifying plaque types and plaque burden prior to stent implantation. The study classified quadrants of IVOCT images as either fibrous, lipid, calcium, or normal. Core Lab analysts were well trained in interpreting OCT image data. For our study, the analyst determined start and stop image frames for a homogeneous VOI. Then, in the view, the analyst segmented the VOI using freehand brush strokes and applied a plaque-type label (fibrous, lipid, or calcified). All segmentations and labels for VOIs were saved in a convenient data structure for easy retrieval and analysis. Standard criteria were used by analysts to identify the various plaque types in pullback image frames.13 Fibrous plaques were characterized by homogeneous, signal-rich regions; calcified regions by signal-poor regions with sharp borders; and lipid plaques by signal-poor regions with diffuse borders. In this study, we used 311 VOIs from 35 pullbacks. The VOIs were of various sizes and shapes. Most consisted of 2 to 5 image frames, 50 to 200 A-lines, and 20 to 50 sample points in each A-line. 3.2.Parameter Estimation of Tissue Optical PropertiesEach VOI was processed as follows. We applied 2-D noise reduction filtering to linear data using a variety of filters [average filter (AF), median filter (MF), and Lee speckle reduction filter (LF)39] with parameters listed later. To avoid any “edge effects” during filtering, we applied 2-D filtering to entire image frames and subsequently used the stored spatial coordinates of each VOI to extract a “filtered” VOI. Any A-line with fewer than five samples (points) was removed from consideration. Rearranging and taking the natural logarithm of Eq. (1), we get the equation below: In the event that the left-side term within square brackets was , we set its value to . Equation (4) reduces to fitting a straight line to the data. Parameter estimation methods included individual line (IL) fits and parallel line (PL) fits to A-lines. In IL fitting, a slope and intercept are fit to each A-line in a collection of adjacent A-lines. In PL fitting, we fit one slope and multiple intercepts to the collection as is rationalized next. Assuming locally homogeneous tissue within a VOI, there should be a single characteristic value and a single slope. Intercepts depend upon distance from the catheter, the angle of incidence with the lumen, depth in tissue, and so on, which are observations consistent with the PL approach. Moreover, a PLs’ model has many fewer parameters making it less sensitive to noise. In addition, we have also employed methods that perform statistical analysis to reject data outliers for which we use the term “robust.” The methods were:
3.3.Data AnalysisFor each VOI, we obtained and estimates for each combination of filtering (three methods) and parameter estimation (eight methods). Goodness-of-fit measures included coefficient of variation (CV) given by the ratio of the parameter uncertainty to the point estimate, the root mean square error (RMSE), and adjusted , calculated as described in Ref. 40. For good filtering and estimation methods, we want: (1) small parameter uncertainty intervals, (2) small spread of point estimates across many VOIs of a particular tissue type, (3) superior goodness-of-fit values (high adjusted , low RMSE, and low CV), and (4) high sensitivity/specificity from parameter estimates using a supervised classifier. 3.4.Processing ParametersWe optimized the processing parameters in preliminary experiments. The size of the filter window () and other parameters was tuned by examining their role on image appearance as well as estimates. It should be noted that a filter kernel of length 7 is approximately the same size as the axial point spread function of our system. We used a window for the LF, AF, and MF. Since VOIs were of irregular shape, we obtained some short A-line segments which were omitted if they contained fewer than five samples. To avoid fitting the baseline noise floor, we ended consideration of A-line samples along when there was a run of nine samples below the baseline value of six in the original linear data. This occurred in less than 1% of A-lines as experts did not mark VOIs at large depths in the tissue. For robust estimation with trimming, we experimentally determined an optimal value of . For iterative estimation methods, we experimentally determined that three iterations and two SDs gave good results. More details about optimization experiments are in Sec. 4. In an exploratory classification experiment, we used three features (, , ), and a support vector machine (SVM) classifier with a linear kernel, fivefold cross validation, and feature normalization. We used one-against-all to identify three tissue types. Parameter settings of the classifier were determined from preliminary optimization experiments with a subset of VOIs. We weighted samples in inverse proportion to their prevalence.41 Performance results were averaged across the five folds. 4.ResultsExample IVOCT images in Figs. 1(a)–1(d) show calcified, fibrotic, lipid, and normal (media) regions in the view. Figures 1(e) and 1(f) show a sample IVOCT image frame from a clinical pullback in both (anatomical) and (polar) views, with a calcified region identified in red. The VOIs were marked on view because all subsequent processings were done on image data. The VOIs were mapped to for presentation (Fig. 2). We have observed that, in general, the tissue types in a region of interest do not change much across successive frames, i.e., they have a high correlation, justifying the use of volumetric analysis in our study (Fig. 2). Fig. 1Appearance of atherosclerotic tissue types in IVOCT images. They are: (a) fibrous, (b) lipid, (c) calcium, and (d) normal (media). The latter includes a zoomed inset to show the typical layered appearance. In (e) and (f), a sample IVOCT image frame is shown in the (anatomical) and (polar) views, respectively. A calcified region is identified in red in both frames. 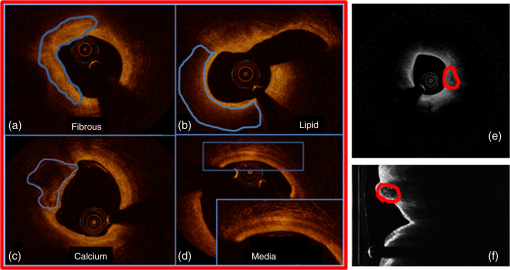 Fig. 2Marking of VOIs in images across frames. Columns contain three successive image frames for (1a–1c) calcium, (2a–2c) lipid, and (3a–3c) fibrous, corresponding to a single VOI. Because tissue types are preserved across a few image frames with spacing, we can process A-lines within VOIs. In practice, such multiple VOIs would be obtained from a single homogenous region (not shown). 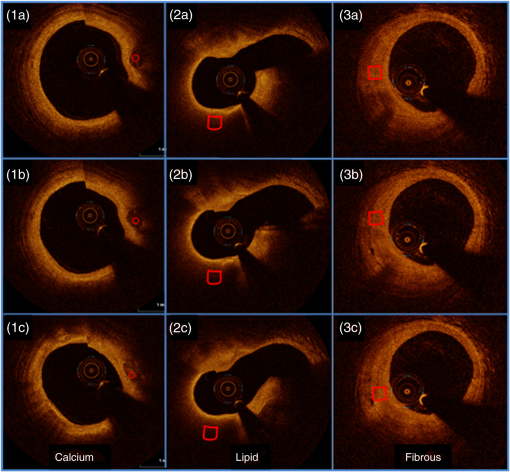 We obtained images of intralipid and water and fitted the data to the system model. Average A-lines from a stationary acquisition are shown in Fig. 3. We ignored the very small upward trend in water and used an average value for . Water images were similar across the three tested catheters, and values were close to baseline values at large in clinical images. Using the average A-line from dilute intralipid images, Eq. (1) was fit to data over the “imaging range” from 1 to 2.5 mm. As described previously, values for , , and were estimated using a least squares fit [Fig. 3(b)] and other parameters were obtained from system specifications. Parameters for the typical catheter in Fig. 3 were: , , , , , and . Fig. 3Determination of imaging system model parameters. The average A-line in water is shown in (a) with intensity on a linear scale. In (b), we plot the average A-line in intralipid ( dilution). We fit data within the “imaging range,” as shown in red. and were estimated from these data. and were otherwise obtained. Image data were acquired in stationary mode and about 100 A-lines across time and were averaged. 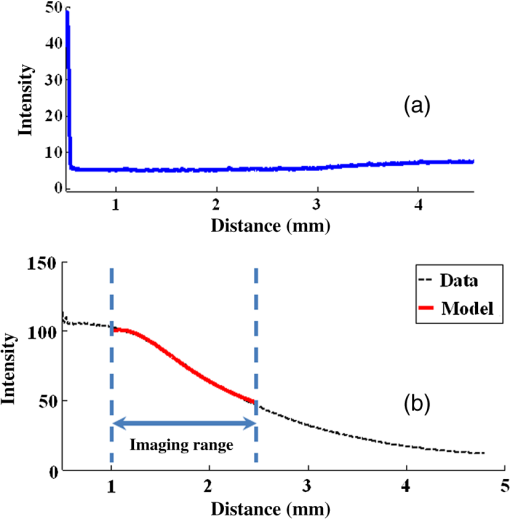 We determined the sensitivity of estimates to the correction model parameter estimates. Table 1 shows the result for representative VOIs using PLs to estimate . Values of changed by only 3% to 7% with and without catheter correction. To study the potential effect of variation between catheters, we perturbed catheter parameters over a range (a variation much larger than observed) and determined the effect on . In general, different catheter correction parameters had a relatively minor effect (typically ) on , smaller than the uncertainty due to noise (see Sec. 4). Varying had a bigger effect than varying . Table 1Effect of catheter correction on μt estimates obtained with the parallel lines’ (PLs’) method for a single-representative VOI from each tissue type. The first two columns show μt estimates within the VOI with and without catheter correction. Fibrotic shows the least variation (3.8%), while calcium (7.8%) and lipid (7.6%) show a larger variation in μt. We also applied a ±5% perturbation to z0 (around a nominal value) keeping zR fixed, and repeated the experiment with a ±5% perturbation to zR (around a nominal value) keeping z0 fixed. Perturbation had a minor effect (<3% in most cases) on μt estimates.
Note: All numbers in table refer to the μt estimate in mm−1. Preprocessing methods were systematically tuned. Individual A-lines with speckle were quite noisy and were visually evident only after filtering trends. Using IL parameter estimation, noise reduction filters substantially reduced SE uncertainties of estimates [Fig. 4(a)]. SE was reduced from 0.8442 [no filtering (NF)] to 0.5371 using AF, 0.446 using MF, and 0.1923 using LF. Therefore, LF was deemed the best filtering approach. Small variations in point estimates were obtained, but effects were inconsistent across VOIs, reducing the concern of potential bias. Effects of noise reduction were less dramatic when using other parameter estimation approaches, but trends were similar. Since we want estimates to help distinguish one tissue type from another, we analyzed the effect of filtering on the spread of estimates for a single tissue type across many VOIs. For calcium, LF reduced the spread of estimates by about 13% [Fig. 4(b)]. Similar results were obtained across other tissue types. Fig. 4Effect of filtering on estimates obtained with IL. In (a), a single calcium VOI with 122 A-lines was analyzed. IL was used to estimate with no filtering (NF) and the three filtering methods—average filter (AF), median filter (MF), and Lee filter (LF). Small () changes of point estimates were obtained, but there were inconsistent trends across other VOIs. Standard errors (SEs) were 0.8442 (NF), 0.5371 (AF), 0.446 (MF), and 0.1923 (LF). NF gave the largest uncertainty SE expressed as a percentage of the point estimates (), while LF gave the smallest (4%). (b) 106 calcium VOIs were analyzed using IL and different filter treatments. Means and standard deviations of are plotted. The LF had the smallest standard deviation across VOIs. 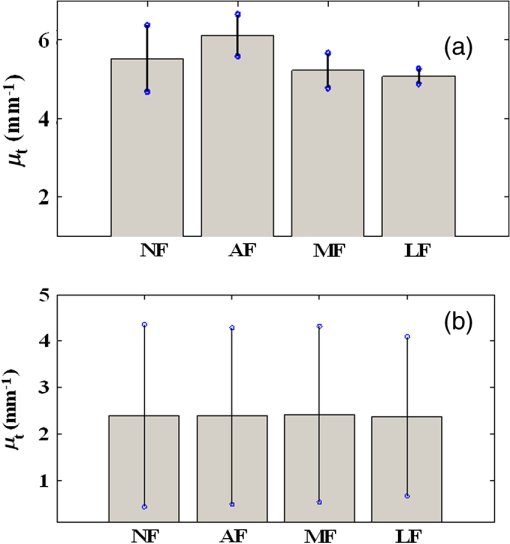 To optimize in noniterative trimming, we varied from 0% to 40% in steps of 5%, and at each step, we computed and the resulting RMSE for each VOI in a set of 84 VOIs drawn from all three plaque types. Figure 5 shows the average RMSE across VOIs plotted against . The optimum RMSE was , and this was used in subsequent experiments. Similarly, we optimized in iterative trimming, which preserved A-lines with in the range (). We estimated and RMSE with , 2, or 3 and obtained an average RMSE versus curve. The optimal was used in subsequent experiments (not shown). Fig. 5Determination of optimal for trimming. Using 84 VOIs drawn from all three plaque types, we obtained RMSE versus for each VOI. The average RMSE curve plotted gave a minimum at . 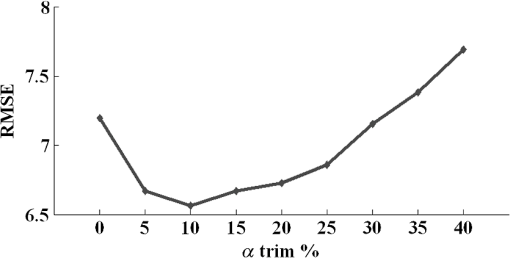 We next compared the parameter estimation approaches. Figure 6 illustrates IL and PL. In this calcified VOI, PL gave a reduced parameter uncertainty (which results in a more robust and accurate estimate) as compared with IL (see legend). Using processing parameters from the above paragraph and LF, we examined the effects from the eight parameter estimation approaches. For a typical calcium VOI, -T-PL and PL gave smaller parameter uncertainty and SEs than other techniques (Fig. 7). There were small changes in point estimates, but across VOIs, there were no obvious trends. Over 111 fibrotic VOIs, the distribution of as a function of estimation methods is shown in Fig. 8(a) without censoring and Fig. 8(b) with censoring. Censoring was done to remove VOIs having bad fits, i.e., . With censoring on RMSE, the number of outliers in was greatly reduced. Since censoring tended to remove VOIs having very large estimates, means were reduced with censoring. Also, the error bars were smaller after censoring as compared with before censoring, which meant that the estimates were more robust after censoring. Fig. 6A subset of seven A-lines from an example calcified VOI (a) is shown. These A-lines are fitted using IL (b) and PL (c). In (a) and (b)s, A-lines have been color coded for clarity. (c) PL fitting where all fitted lines (red) have same slope but different intercepts. The estimates ( error) were IL () and PL (). The PL typically results in smaller parameter uncertainty and a more accurate estimate. 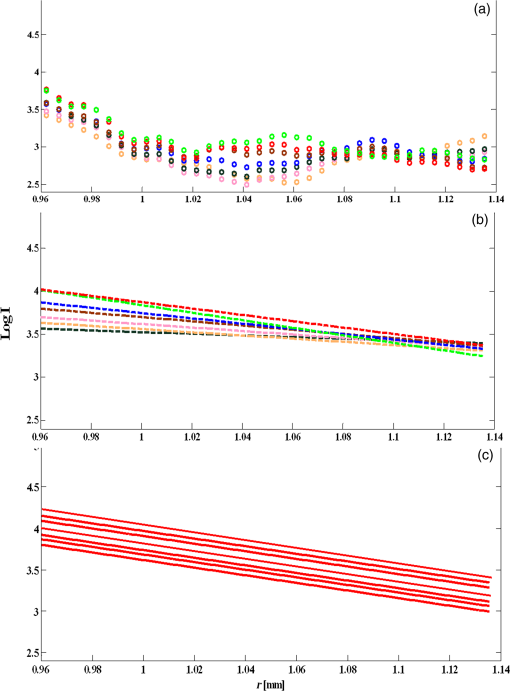 Fig. 7Comparison of parameter estimation methods. A typical calcium VOI containing 62 A-lines was analyzed following LF. The following results were obtained ( error of ): (1) IL: (8 lines removed), (2) -T-IL: (24 lines removed), (3) IT-IL: (14 lines removed), (4) PL: (8 lines removed), (5) -T-PL: (24 lines removed), (6) IT-PL: (14 lines removed), (7) NIL-PL-TM: (38 lines removed), and (8) IL-PL-TM: (39 lines removed). -T-PL and PL methods gave the smallest estimated uncertainty. 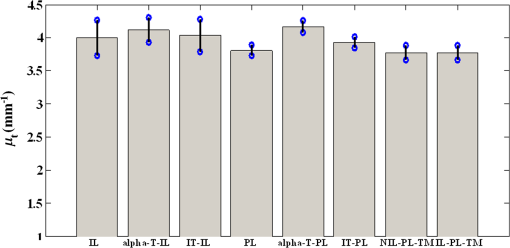 Fig. 8Box whisker plot of as a function of estimation methods for (a) 111 fibrotic VOIs, and (b) a censored subset of 79 VOIs obtained by applying an RMSE threshold of 6. The red bar is the median, the edges of box are 25th and 75th percentiles, and the whiskers represent the most extreme points that are not outliers. Outliers are denoted with red +’s. The RMSE censoring greatly reduced the number of very large outliers and spread of estimates. With censoring, rank ordering on standard deviation across VOIs is . The () of for methods is: IL (), -T-IL (), IT-IL (), PL (), -T-PL (), IT-PL (), NIL-PL-TM (), and IL-PL-TM (). 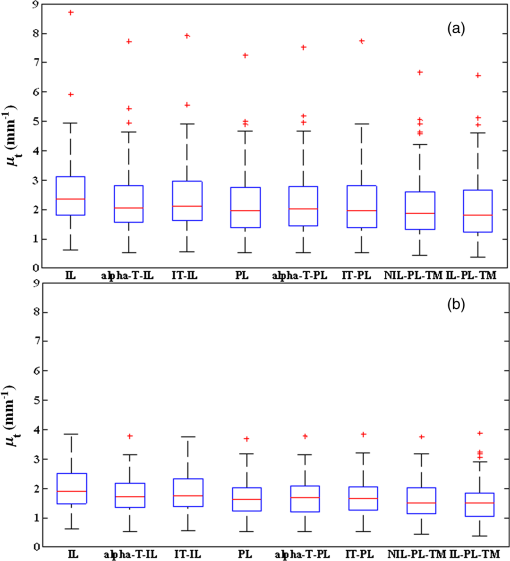 Since we are ultimately interested in automated processing, we analyzed the role of the size of the region of support on estimates. Figure 9 shows a scatter plot of numbers of A-lines in VOIs versus SE of . In general, lower SEs were obtained for VOIs containing more A-lines. A large percentage () of VOIs had 50 or more A-lines in them. This suggests that the experts identified many homogeneous regions with A-lines and average length of 35 samples. The VOIs having at least 50 A-lines resulted in an SE of typically of the estimate. Fig. 9Effect of number of A-lines on estimates. SE of estimates tends to reduce with VOIs having more A-lines. Processing included LF, -T-PL, no censoring. Tissue types are color coded. The mean number of A-lines was 150 and 80% of homogenous regions have A-lines. The solid black line is a linear fit that predicts an uncertainty with 50 A-lines. 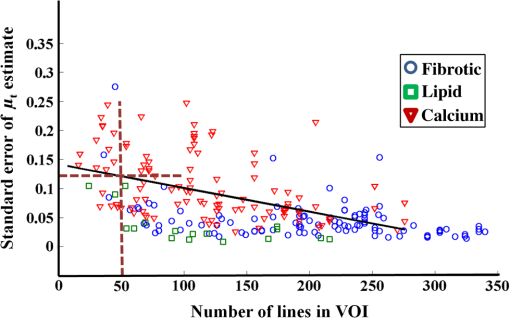 We are especially interested in goodness-of-fit measures because they could become features in an automated machine learning approach to classify plaque types. Adjusted (Ref. 40) is a good method for comparing fits of models with different numbers of parameters [Fig. 10(a)]. Methods -T-PL, PL, and IT-PL had higher adjusted than other methods, indicating the appropriateness of PLs, a general observation across VOIs. We observed improvements with PLs because the number of free parameters was reduced from to , where was the number of A-lines. Although the adjusted is a good method to compare model order, its dependence on slope does not allow comparisons across VOIs. For fibrous VOIs, RMSE and CV histograms were plotted [Figs. 10(b) and 10(c)]. Fig. 10Goodness-of-fit measures. (a) Adjusted is plotted as a function of parameter estimation method for a single-fibrotic VOI with 240 A-lines. The PL, -T-PL, and IT-PL gave higher adjusted than others. (b) Histogram of RMSE for 111 fibrotic VOIs for -T-PL. More than 90% had good fits with . (c) Using the same data, over 85% of VOIs have a . 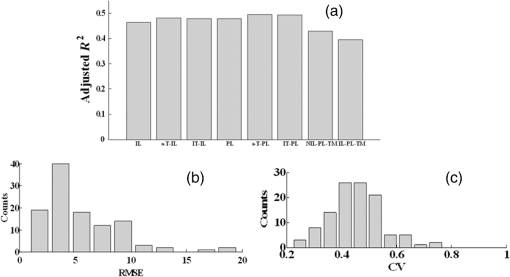 More than 90% of VOIs had . The RMSE was deemed the most appropriate goodness-of-fit for censoring VOIs. When VOIs with large RMSEs were omitted, the variability of estimates was reduced (Fig. 8) and classification (Table 3) improved. To evaluate the usefulness of optical properties as features in an automatic classifier for atherosclerotic plaque tissue types, we used three features (, , ), where was the mean intensity in the VOI. Intensity features are related to the backscattering characteristics of the tissue within the VOI as well as any attenuation prior to the VOI, features which should aid prediction of class membership. We used supervised classification, SVM,41 fivefold cross validation, and one against all for each of the three classes (Table 3). Data were processed without and with RMSE-based censoring, where VOIs with were excluded. Classification improved with censoring [compare Tables 3(a) and 3(b)]. (The RMSE threshold is different from that used in Fig. 8 because we wanted to be uniform across all three tissue classes.) Figure 11(a) shows a clear separation of classes in 3-D feature space following VOI censoring. Features (, ), the two best features, also demonstrated good separation in Fig. 11(b). Fig. 11(, , ) were used as features in automatic classification of VOIs using an SVM classifier into three plaque types—fibrotic, lipid, and calcium. All three features are plotted in (a) and the two best features are plotted in (b). 311 expert annotated VOIs from 35 IVOCT pullbacks. We used -T-PL, LF, and censoring of VOIs with , leaving 277. Feature space indicates a good separation between all three plaque types (calcified, fibrotic, and lipid). The () of estimates was: calcified (), fibrotic (), and lipid (). Classification results are in Table 3. 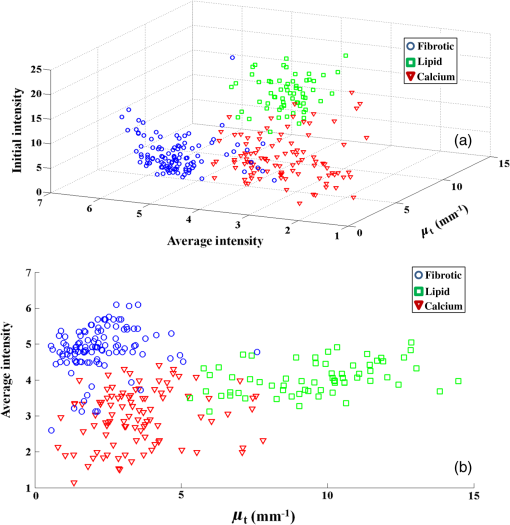 With all analyses considered, we determined that the PLs and trimming were both useful, with PL and -T-PL producing numerically close estimates. In general, we recommend -T-PL, LF, and possible censoring on RMSE value as best choices. Using these choices, we obtained the following values of : calcified (), fibrotic (), and lipid (). Computational estimation of allowed us to distinguish between lipid plaques and calcium plaques with a diffuse edge (Fig. 12). Typically, this is resolved by experts only after analyzing adjacent frames. Fig. 12Our robust estimation methods have helped us distinguish between a lipid plaque (a) and a calcified plaque with a diffuse edge (b), both shown by blue arcs. This is often a source of confusion at first glance during manual analysis. When we employed parameter estimation of , we could see a significant numerical difference between the two cases (lipid: , calcium with diffuse edge: ), which we believe would aid in proper classification. 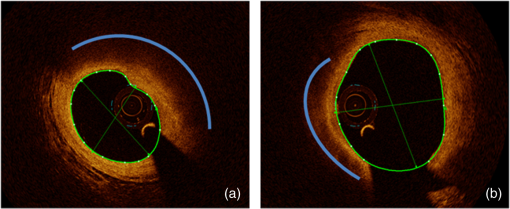 5.DiscussionThe characterization of tissue optical properties from 3-D IVOCT clinical pullbacks will be an important step in reliable tissue classification. Previous methods for estimating optical properties have typically used time-averaged data in a stationary acquisition29 or single A-lines from a pullback.30 Since stationary acquisitions are clinically impractical and single A-lines are extremely noisy, we created a 3-D approach whereby groups of A-lines close to each other are analyzed together. Innovations in this report include: use of A-lines from within a 3-D VOI to improve stability of estimates; advanced, robust parameter estimation approaches including goodness-of-fit estimates; and a rigorous methodology for comparing estimation methods applied to clinical pullback data. Results suggest that filtering is desirable, that estimates vary some with the method used, that PL and its variants generally outperformed IL and its variants, and that RMSE is the favored goodness-of-fit measure which can be used effectively to “censor” VOIs with bad fits. As detailed later, we deemed -T-PL as best. In general, we can conclude that the PL model, which fits one slope and multiple intercepts to a collection of spatially adjacent A-lines, is appropriate since there should be a single characteristic value and a single slope for locally homogeneous tissue. LF39 was the best filtering method among those tested. Filtering visually reduced the noise in A-lines. Comparing filters on a single VOI, the uncertainty of was minimized with the LF [Fig. 4(a)]. Also, LF reduced the spread of across many VOIs for a given tissue [Fig. 4(b)], presumably enabling better separation of tissues. We did not observe any consistent bias due to filtering on estimates. It is satisfying that a filter designed for speckle noise gave the best results for parameter estimation because speckle is the dominant noise in OCT images. Comparing parameter estimation methods via multiple assessments, we determined that PLs and trimming were useful and that the best overall method was -T-PL. Comparing results on a typical VOI, the uncertainty of was least for -T-PL (Fig. 7). Since we are ultimately interested in separating tissue types, reducing the spread of estimates across many VOIs is desirable. -T-PL and PL gave a reduced spread of across many VOIs as compared with other methods (Fig. 8, legend). Since results were variable across filters, VOIs, tissue types, and estimation methods, we developed a voting scheme for comparing parameter estimation methods. For each VOI, we picked a winning technique based on two different criteria: (1) the smallest uncertainty SE of estimate, and (2) the smallest RMSE goodness-of-fit for a VOI. We then counted across all VOIs the number of times each technique was a winner and sorted results. Based on criterion (1), we obtained the following ranking of the four best techniques: . Based on (2), we obtained the following ranking: . Since -T-PL was in the top two for both criteria, we deemed it best. Although IL-PL-TM had the lowest RMSE, it had a large SE uncertainty [it was not in the top four based on criterion (1)]. It is worth noting that having a smaller SE uncertainty is more significant. This is because sometimes a good numerical fit could be obtained with a high SE. Similarly, -T-PL was voted best when we rank-ordered standard deviations across many VOIs of each tissue type. The computational complexity of -T-PL can be greatly reduced by saving intermediate calculations. There have been previous reports of (total attenuation coefficient) and (scattering coefficient) for atherosclerotic tissues in the literature.29–33 Our estimates of are consistent with two previously reported studies that have estimated optical properties of clinically obtained atherosclerotic plaques using the same imaging system but different estimation procedures on stationary IVOCT acquisitions29,30 (Table 2) [calcified (ours: , reported: 2 to ), fibrotic (ours: , reported: 2 to ), and lipid (ours: , reported: )]. Some other studies32,33 report much lower values of for lipid ( and ) and much higher values for calcified (), but these studies used ex vivo images, a bench-top OCT scanner, and a different light source, therefore, they are not comparable to our study. Another study at 31 reported and values instead of making comparisons difficult. Table 2Estimates of μt are consistent with values from two previously reported studies29,30 that have estimated optical properties of atherosclerotic plaques using the same imaging system (1310-nm center wavelength and 110-nm wavelength range).
The robust, 3-D methods proposed here for measuring optical properties of tissue should be applicable to other OCT applications. Specifically, methods should be applicable to volumetric OCT cancer imaging, where optical properties have been used to distinguish cancerous tissues.42–44 Because it is desirable to include the quality of assessment with parameter estimates, we investigated goodness-of-fit measures. Adjusted is a good way to compare models of different orders. The PL methods (-T-PL, PL, and IT-PL) gave higher adjusted than other methods (Fig. 10), but depends on slope and is inappropriate for assessing the fit in a particular VOI. The CV was considered, but it was determined that the CV simply favored VOIs with more A-lines. We prefer RMSE because it is simple to compute and interpret. A histogram of RMSE across many VOIs [Fig. 10(b)] shows that 90% of VOIs have “good fits” with . Classification results were improved when VOIs with were censored, indicating that the RMSE could be a useful feature for automated processing going forward. For example, one could estimate from regions, but additionally use the computed RMSE value as a confidence in the estimate and weight the feature accordingly during classification. The exploratory classification study yielded surprisingly good results based solely on optical property parameters (Table 3, Fig. 11). The choice of classifier and other parameter settings was determined from preliminary optimization experiments in which a small subset of VOIs of all three plaque types was employed. We used SVM, three tissue types, one-against-all training, three features (, , ), and fivefold cross validation to assess results. Censoring VOI data with poor fits () improved sensitivity and specificity as compared with uncensored data for calcium and lipid VOIs, and only slightly lowered specificity (while keeping the same sensitivity) for fibrotic VOIs [Tables 3(a) versus 3(b)]. It was encouraging that the optical parameter estimates allowed us to distinguish between lipid plaques and calcium plaques with a diffuse edge, often a problem during manual analysis (Fig. 12). In manual analysis, analysts typically analyze adjacent 2-D frames to make a determination using standard definitions provided by Yabushita et al.13 It is promising that the computational methods can determine this from optical properties alone. Table 3Classifier performance using optical properties features (μt, 〈I〉, I0). (a) Classifier performance on 311 VOIs using α-T-PL, SVM, one-versus-rest, and fivefold cross validation. (b) VOIs with RMSE>12 were removed, leaving 277 VOIs. Censoring improved classifier performance, particularly for lipid.
The aim of our study was to compare advanced parameter estimation methods applicable to clinical 3-D IVOCT pullbacks. This avoids the use of ex vivo data with its potential limitations including potential tissue degradation, no mean or pulsating pressure, no cardiac motion, unrealistic temperatures, no blood artifacts, and a straight path pullback rather than actual 3-D coronary geometry. Moreover, in the event of spatially inaccurate, sparse histological sampling, one can make inappropriate “ground truth” determinations using ex vivo data. Instead we used as ground truth VOIs identified by well-trained analysts using well-established criteria. Our VOI classification results overestimate that which would be obtained in automatic processing of pullback data. That is, analysts choose homogeneous, easily identified VOIs, giving rise to relatively unambiguous computer classification. In addition, analysts purposely did not choose regions of macrophages and thrombi because we were focusing on optical attenuation estimation rather than other features. Our approach should be quite appropriate for optimizing the 3-D parameter estimation methods in this report. In summary, we believe that the robust processing methods make it possible to assess tissue optical properties from noisy, IVOCT clinical pullback data. In addition, optical parameters thus obtained will be important image features for an automated analysis of plaques. We believe that the 3-D methods reported here for assessing optical properties will also be applicable in volumetric OCT cancer imaging. AcknowledgmentsThis project was supported by the National Heart, Lung, and Blood Institute through Grants NIH R21HL108263 and 1R01HL114406-01 and by the National Center for Research Resources and the National Center for Advancing Translational Sciences through Grant UL1RR024989. These grants are a collaboration between Case Western Reserve University and University Hospitals of Cleveland. ReferencesI. K. Janget al.,
“In vivo characterization of coronary atherosclerotic plaque by use of optical coherence tomography,”
Circulation, 111 1551
–1555
(2005). http://dx.doi.org/10.1161/01.CIR.0000159354.43778.69 CIRCAZ 0009-7322 Google Scholar
T. Kumeet al.,
“Assessment of coronary arterial plaque by optical coherence tomography,”
Am. J. Cardiol., 97 1172
–1175
(2006). http://dx.doi.org/10.1016/j.amjcard.2005.11.035 AJNCE4 0258-4425 Google Scholar
F. Pratiet al.,
“Expert review document on methodology, terminology, and clinical applications of optical coherence tomography: physical principles, methodology of image acquisition, and clinical application for assessment of coronary arteries and atherosclerosis,”
Eur. Heart J., 31 401
–415
(2010). http://dx.doi.org/10.1093/eurheartj/ehp433 EHJODF 0195-668X Google Scholar
R. PuriM. I. WorthleyS. J. Nicholls,
“Intravascular imaging of vulnerable coronary plaque: current and future concepts,”
Nat. Rev. Cardiol., 8 131
–139
(2011). http://dx.doi.org/10.1038/nrcardio.2010.210 NRCAE6 1759-5002 Google Scholar
D. StamperN. J. WeissmanM. Brezinski,
“Plaque characterization with optical coherence tomography,”
J. Am. Coll. Cardiol., 47 C69
–C79
(2006). http://dx.doi.org/10.1016/j.jacc.2005.10.067 JACCDI 0735-1097 Google Scholar
W. M. Suhet al.,
“Intravascular detection of the vulnerable plaque,”
Circ.: Cardiovasc. Imaging, 4
(2), 169
–178
(2011). http://dx.doi.org/10.1161/CIRCIMAGING.110.958777 Google Scholar
M. J. Suteret al.,
“Progress in intracoronary optical coherence tomography,”
IEEE J. Sel. Top. Quantum Electron., 16 706
–714
(2010). http://dx.doi.org/10.1109/JSTQE.2009.2035333 IJSQEN 1077-260X Google Scholar
G. J. TearneyI. K. JangB. E. Bouma,
“Imaging coronary atherosclerosis and vulnerable plaques with optical coherence tomography,”
Optical Coherence Tomography, 1083
–1101 Springer, Berlin, Heidelberg
(2008). Google Scholar
G. J. Tearneyet al.,
“Quantification of macrophage content in atherosclerotic plaques by optical coherence tomography,”
Circulation, 107 113
–119
(2003). http://dx.doi.org/10.1161/01.CIR.0000044384.41037.43 CIRCAZ 0009-7322 Google Scholar
G. J. TearneyI. K. JangB. E. Bouma,
“Optical coherence tomography for imaging the vulnerable plaque,”
J. Biomed. Opt., 11 021002
(2006). http://dx.doi.org/10.1117/1.2192697 JBOPFO 1083-3668 Google Scholar
S. Taharaet al.,
“Intravascular optical coherence tomography detection of atherosclerosis and inflammation in murine aorta,”
Arterioscler., Thromb., Vasc. Biol., 32 1150
–1157
(2012). http://dx.doi.org/10.1161/ATVBAHA.111.243626 ATVBFA 1079-5642 Google Scholar
H. G. Bezerraet al.,
“Intracoronary optical coherence tomography: a comprehensive review, clinical and research applications,”
JACC: Cardiovasc. Interventions, 2 1035
–1046
(2009). http://dx.doi.org/10.1016/j.jcin.2009.06.019 1936-8798 Google Scholar
H. Yabushitaet al.,
“Characterization of human atherosclerosis by optical coherence tomography,”
Circulation, 106 1640
–1645
(2002). http://dx.doi.org/10.1161/01.CIR.0000029927.92825.F6 CIRCAZ 0009-7322 Google Scholar
I. K. Janget al.,
“Visualization of coronary atherosclerotic plaques in patients using optical coherence tomography: comparison with intravascular ultrasound,”
J. Am. Coll. Cardiol., 39 604
–609
(2002). http://dx.doi.org/10.1016/S0735-1097(01)01799-5 JACCDI 0735-1097 Google Scholar
Z. Wanget al.,
“Semiautomatic segmentation and quantification of calcified plaques in intracoronary optical coherence tomography images,”
J. Biomed. Opt., 15 061711
(2010). http://dx.doi.org/10.1117/1.3506212 JBOPFO 1083-3668 Google Scholar
Z. Wanget al.,
“Automatic segmentation of intravascular optical coherence tomography images for facilitating quantitative diagnosis of atherosclerosis,”
Proc. SPIE, 7889 78890N
(2011). http://dx.doi.org/10.1117/12.876003 PSISDG 0277-786X Google Scholar
Z. Wanget al.,
“Volumetric quantification of fibrous caps using intravascular optical coherence tomography,”
Biomed. Opt. Express, 3 1413
(2012). http://dx.doi.org/10.1364/BOE.3.001413 BOEICL 2156-7085 Google Scholar
Z. Wanget al.,
“Single-shot stent segmentation in intravascular OCT pullbacks,”
in Biomedical Optics,
BTu4B-5
(2012). Google Scholar
H. Luet al.,
“Automatic stent strut detection in intravascular OCT images using image processing and classification technique,”
Proc. SPIE, 8670 867015
(2013). http://dx.doi.org/10.1117/12.2007183 PSISDG 0277-786X Google Scholar
H. Luet al.,
“Automatic stent detection in intravascular OCT images using bagged decision trees,”
Biomed. Opt. Express, 3 2809
–2824
(2012). http://dx.doi.org/10.1364/BOE.3.002809 BOEICL 2156-7085 Google Scholar
S. Taharaet al.,
“Angiographic, IVUS and OCT evaluation of the long-term impact of coronary disease severity at the site of overlapping drug-eluting and bare metal stents: a substudy of the ODESSA trial,”
Heart, 96 1574
–1578
(2010). http://dx.doi.org/10.1136/hrt.2009.188037 1355-6037 Google Scholar
H. Kyonoet al.,
“Optical coherence tomography (OCT) strut-level analysis of drug-eluting stents (DES) in human coronary bifurcations,”
EuroIntervention, 6 69
–77
(2010). http://dx.doi.org/10.4244/EIJV6I1A11 1774-024X Google Scholar
G. Guagliumiet al.,
“Strut coverage and vessel wall response to a new-generation paclitaxel-eluting stent with an ultrathin biodegradable abluminal polymer: Optical Coherence Tomography Drug-Eluting Stent Investigation (OCTDESI),”
Circ. Cardiovasc. Intervention, 3
(4), 367
–375
(2010). http://dx.doi.org/10.1161/CIRCINTERVENTIONS.110.950154 Google Scholar
G. Guagliumiet al.,
“Optical coherence tomography assessment of in vivo vascular response after implantation of overlapping bare-metal and drug-eluting stents,”
JACC: Cardiovasc. Interventions, 3 531
–539
(2010). http://dx.doi.org/10.1016/j.jcin.2010.02.008 1936-8798 Google Scholar
S. J. Kanget al.,
“Optical coherence tomographic analysis of in-stent neoatherosclerosis after drug-eluting stent implantation,”
Circulation, 123 2954
–2963
(2011). http://dx.doi.org/10.1161/CIRCULATIONAHA.110.988436 CIRCAZ 0009-7322 Google Scholar
A. KnuS. BonevW. Knaak,
“New method for evaluation of in vivo scattering and refractive index properties obtained with optical coherence tomography,”
J. Biomed. Opt., 9 265
–273
(2004). http://dx.doi.org/10.1117/1.1647544 JBOPFO 1083-3668 Google Scholar
O. Manfriniet al.,
“Sources of error and interpretation of plaque morphology by optical coherence tomography,”
Am. J. Cardiol., 98 156
–159
(2006). http://dx.doi.org/10.1016/j.amjcard.2006.01.097 AJNCE4 0258-4425 Google Scholar
G. van Soestet al.,
“Pitfalls in plaque characterization by OCT image artifacts in native coronary arteries,”
JACC Cardiovasc. Imaging, 4 810
–813
(2011). http://dx.doi.org/10.1016/j.jcmg.2011.01.022 1936-8798 Google Scholar
C. Xuet al.,
“Characterization of atherosclerosis plaques by measuring both backscattering and attenuation coefficients in optical coherence tomography,”
J. Biomed. Opt., 13 034003
(2008). http://dx.doi.org/10.1117/1.2927464 JBOPFO 1083-3668 Google Scholar
G. van Soestet al.,
“Atherosclerotic tissue characterization in vivo by optical coherence tomography attenuation imaging,”
J. Biomed. Opt., 15 011105
(2010). http://dx.doi.org/10.1117/1.3280271 JBOPFO 1083-3668 Google Scholar
D. Levitzet al.,
“Determination of optical scattering properties of highly-scattering media in optical coherence tomography images,”
Opt. Express, 12 249
–259
(2004). http://dx.doi.org/10.1364/OPEX.12.000249 OPEXFF 1094-4087 Google Scholar
D. J. Faberet al.,
“Quantitative measurement of attenuation coefficients of weakly scattering media using optical coherence tomography,”
Opt. Express, 12 4353
–4365
(2004). http://dx.doi.org/10.1364/OPEX.12.004353 OPEXFF 1094-4087 Google Scholar
F. J. Van der Meeret al.,
“Localized measurement of optical attenuation coefficients of atherosclerotic plaque constituents by quantitative optical coherence tomography,”
IEEE Trans. Med. Imaging, 24 1369
–1376
(2005). http://dx.doi.org/10.1109/TMI.2005.854297 ITMID4 0278-0062 Google Scholar
G. van Soestet al.,
“Frequency domain multiplexing for speckle reduction in optical coherence tomography,”
J. Biomed. Opt., 17 076018
(2012). http://dx.doi.org/10.1117/1.JBO.17.7.076018 JBOPFO 1083-3668 Google Scholar
Y. Yanget al.,
“Optical scattering coefficient estimated by optical coherence tomography correlates with collagen content in ovarian tissue,”
J. Biomed. Opt., 16 090504
(2011). http://dx.doi.org/10.1117/1.3625247 JBOPFO 1083-3668 Google Scholar
P. LeeW. GaoX. Zhang,
“Performance of single-scattering model versus multiple-scattering model in the determination of optical properties of biological tissue with optical coherence tomography,”
Appl. Opt., 49 3538
–3544
(2010). http://dx.doi.org/10.1364/AO.49.003538 APOPAI 0003-6935 Google Scholar
L. S. Athanasiouet al.,
“Methodology for fully automated segmentation and plaque characterization in intracoronary optical coherence tomography images,”
J. Biomed. Opt., 19 026009
(2014). http://dx.doi.org/10.1117/1.JBO.19.2.026009 JBOPFO 1083-3668 Google Scholar
G. J. Ughiet al.,
“Automated tissue characterization of in vivo atherosclerotic plaques by intravascular optical coherence tomography images,”
Biomed. Opt. Express, 4 1014
–1030
(2013). http://dx.doi.org/10.1364/BOE.4.001014 BOEICL 2156-7085 Google Scholar
J. S. Lee,
“Speckle analysis and smoothing of synthetic aperture radar images,”
Comput. Graphics Image Process., 17 24
–32
(1981). http://dx.doi.org/10.1016/S0146-664X(81)80005-6 CGIPBG 0146-664X Google Scholar
H. Theil, Economic Forecasts and Policy, North-Holland Publication(1961). Google Scholar
Y. Tanget al.,
“SVMs modeling for highly imbalanced classification,”
IEEE Trans. Syst., Man, Cybern. B, 39 281
–288
(2009). http://dx.doi.org/10.1109/TSMCB.2008.2002909 ITSCFI 1083-4419 Google Scholar
S. Wanget al.,
“Three-dimensional computational analysis of optical coherence tomography images for the detection of soft tissue sarcomas,”
J. Biomed. Opt., 19
(2), 021102
(2014). http://dx.doi.org/10.1117/1.JBO.19.2.021102 JBOPFO 1083-3668 Google Scholar
L. Scolaroet al.,
“Parametric imaging of the local attenuation coefficient in human axillary lymph nodes assessed using optical coherence tomography,”
Biomed. Opt. Express, 3
(2), 366
–379
(2012). http://dx.doi.org/10.1364/BOE.3.000366 BOEICL 2156-7085 Google Scholar
P. Robbinset al.,
“Parametric imaging of cancer with optical coherence tomography,”
J. Biomed. Opt., 15
(4), 046029
(2010). http://dx.doi.org/10.1117/1.3479931 JBOPFO 1083-3668 Google Scholar
BiographyMadhusudhana Gargesha is a senior research associate in the Department of Biomedical Engineering at Case Western Reserve University, Cleveland, Ohio, USA. He received his BE degree in electronics and communication engineering from Bangalore University, Bangalore, India, in 1997, and his MS and PhD degrees in electrical engineering from Arizona State University, Tempe, Arizona, in 2002 and 2006, respectively. He is the author of more than 15 journal papers and 20 conference publications. His current research interests include 3-D image analysis, processing, and visualization as applied to medical and biological imaging modalities including IVOCT, MRI, and block-face cryo-imaging. Ronny Shalev spent much of the past 21 years in executive positions in public and private companies. Currently, he is an advanced PhD student having extraordinary product development experience. He holds an MSc in electrical engineering and applied physics. His current research interests include medical image processing, machine learning, and visualization as applied to various medical and biological imaging modalities. David Prabhu is a senior level PhD candidate at Case Western Reserve University. He is a Choose Ohio First Scholar and has industrial experience in the field of cryo-imaging. His research interests include image processing and analysis, 3-D visualization, histology, and cryo-imaging. Kentaro Tanaka is the research fellow of the Cardiovascular Imaging Core Laboratory, Harrington Heart and Vascular Institute, University Hospitals Case Medical Center. His interests are cardiovascular imaging, coronary artery OCT, and IVUS. Andrew M. Rollins is a professor of biomedical engineering and medicine at Case Western Reserve University. His research interests are in the development and application of advanced optical technologies, especially optical coherence tomography and optical stimulation, to the study of developmental biology and to endoscopic imaging of cardiovascular disease and cancer. He directs an NIH-funded research program and has trained numerous MS and PhD students. Marco Costa is a professor of medicine, CWRU, Angela & James Hambrick Master Clinician in Innovation, board-certified interventional cardiologist, director of the Interventional Cardiovascular Center, and director of the Research and Innovation Center University Hospitals, Case Medical Center. He has been a leader in developing IVOCT as well as new interventional procedures. He was the first person in the US to perform clinical OCT imaging with an FDA-approved OCT imaging system from LightLab, now St. Jude. Hiram G. Bezerra is an assistant professor of medicine, CWRU, board-certified interventional cardiologist, and medical director of the Cardiovascular Imaging Core Laboratory at University Hospitals (Core Lab), Case Medical Center. Under his Core Lab duties, he is responsible for the analysis of thousands of cardiovascular images from IVOCT as well as other modalities, e.g., intravascular ultrasound, quantitative angiography, cardiac MRI, and cardiac CT. He directs both state and federal research grants. David L. Wilson is the Robert Herbold professor of biomedical engineering and radiology, Case Western Reserve University. His interests are image processing and analysis, cardiovascular imaging, coronary artery OCT, and cryo-imaging. He serves on NIH study sections, editorial boards, and conference committees. He has trained numerous PhD students and postdocs, all of whom are quite exceptional. In addition to federal and state funded research activities, he is the director of CWRU’s Interdisciplinary Biomedical Imaging NIH T32 training grant. |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||

